Age prediction using Microbiota
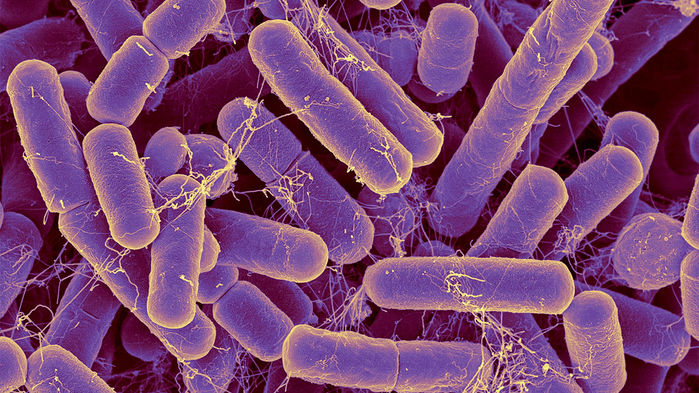
Bacteroides are the most common bacteria species found in the human intestinal tract. Dennis Kunkel Microscopy/Science Source
The idea that you can predict someone’s age based on their gut microbiome is “very plausible” and of “tremendous interest” to scientists.
Dr. Zhavoronkov, longevity researcher at InSilico Medicine, Maryland and computer scientist and microbiome researcher, Robin Knight, director of the Center for Microbiome Innovation at the University of California, San Diego, and colleagues found out how the microbiome changes over time by examining more than 13,000 samples of gut bacteria from healthy individuals living across the globe.
Please see below Dr. Alex Zhavoronkov and colleagues's abstract:
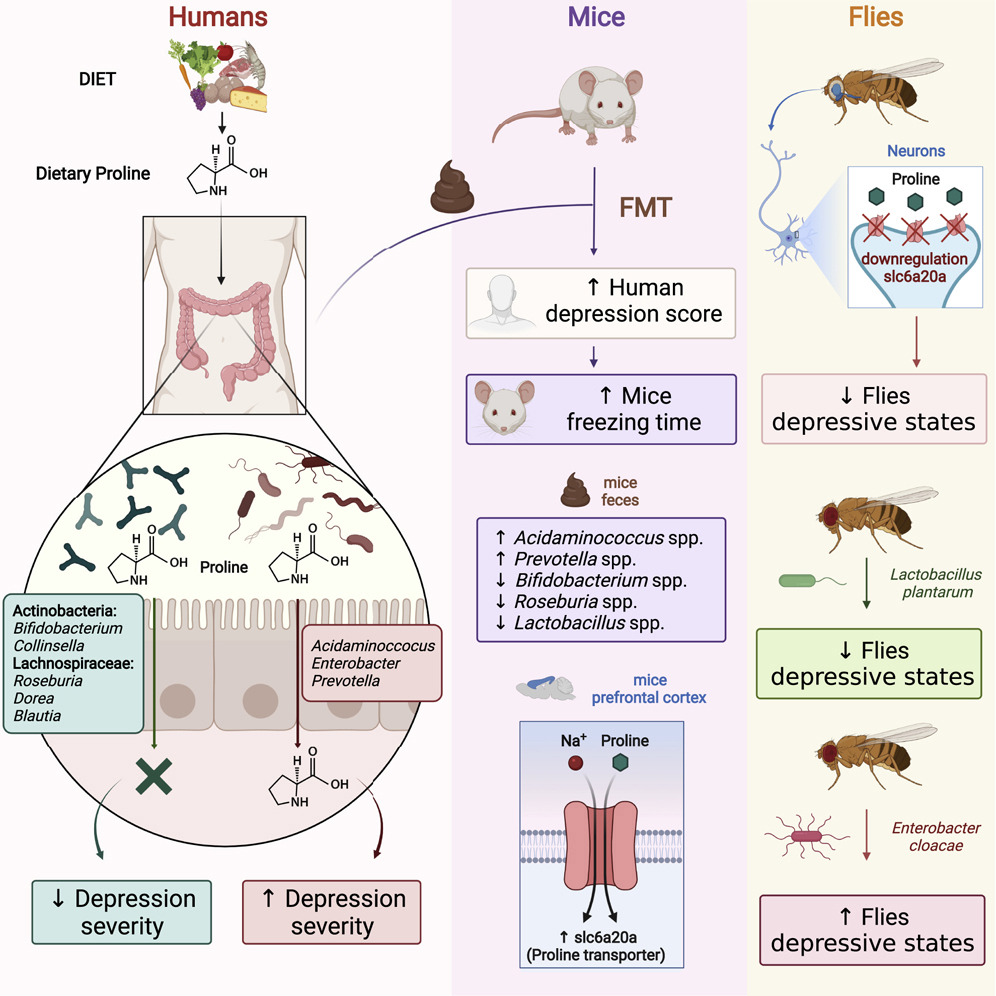
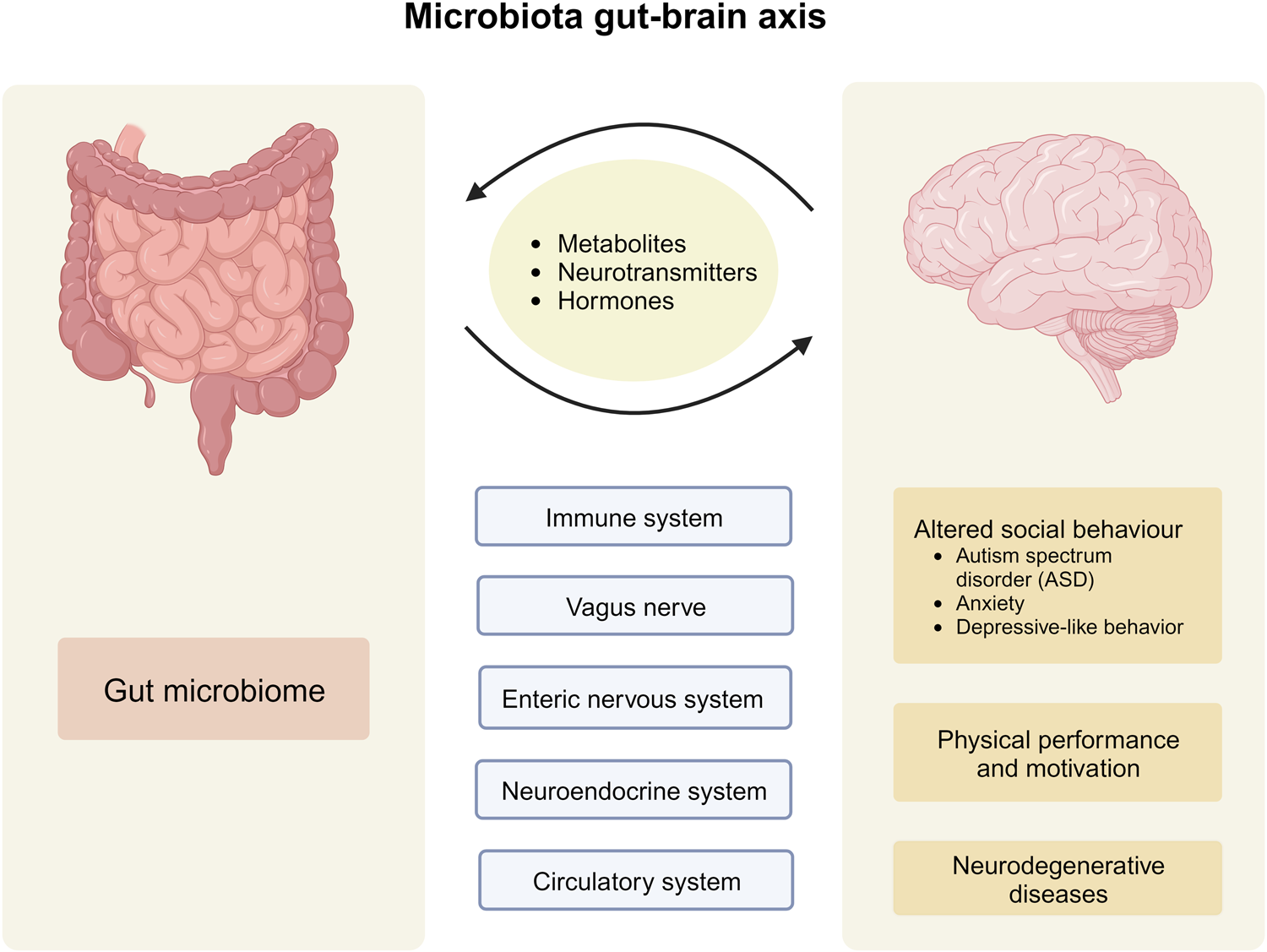
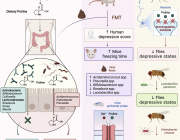
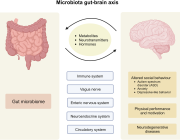
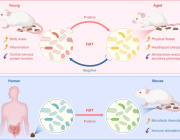
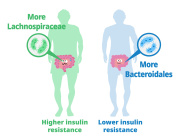
The human gut microbiome is a complex ecosystem that both affects and is affected by its host status. Zhavoronkov says this “microbiome aging clock” could be used as a baseline to test how fast or slow a person’s gut is aging
Previous analyses of gut microflora revealed associations between specific microbes and host health and disease status, genotype and diet.
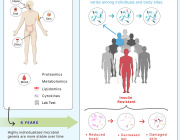
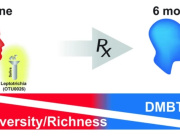
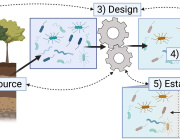
Here, we developed a method of predicting biological age of the host based on the microbiological profiles of gut microbiota using a curated dataset of 1,165 healthy individuals (3,663 microbiome samples).
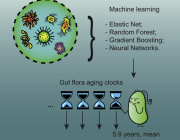
Our predictive model, a human microbiome clock, has an architecture of a deep neural network and achieves the accuracy of 3.94 years mean absolute error in cross-validation.
The performance of the deep microbiome clock was also evaluated on several additional populations.
We further introduce a platform for biological interpretation of individual microbial features used in age models, which relies on permutation feature importance and accumulated local effects.
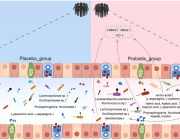
This approach has allowed us to define two lists of 95 intestinal biomarkers of human aging. We further show that this list can be reduced to 39 taxa that convey the most information on their host’s aging.
Overall, we show that (a) micro biological profiles can be used to predict human age; and (b) microbial features selected by models are age-related.
These topics will be discussed during the conference, in the session Microbiota and recent scientific advances.
https://www.biorxiv.org/content/early/2018/12/28/507780