Reconstruction of Ancient Microbial Genomes from the Human Gut
A new study reports on Reconstruction of ancient microbial genomes from the human gut, published by Aleksandar D. Kostic and al.

Image By Danielle Masterson, Microgen
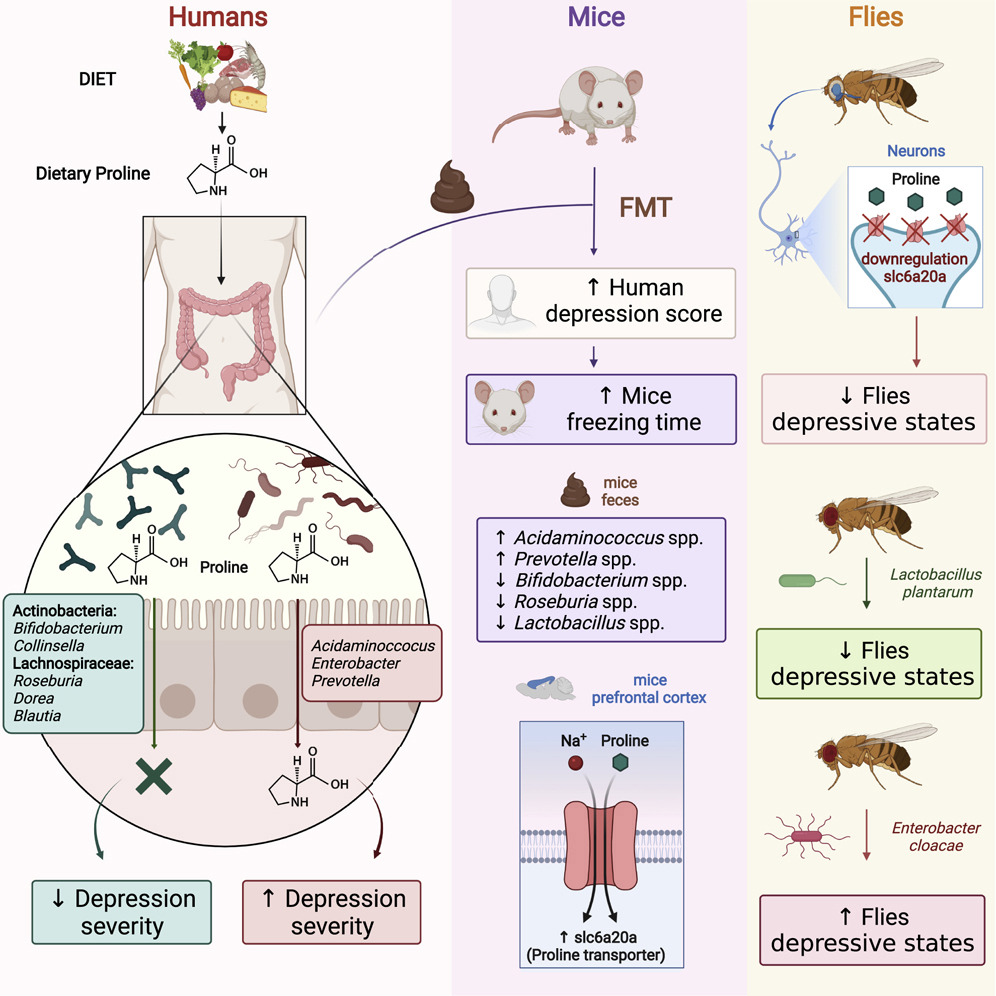
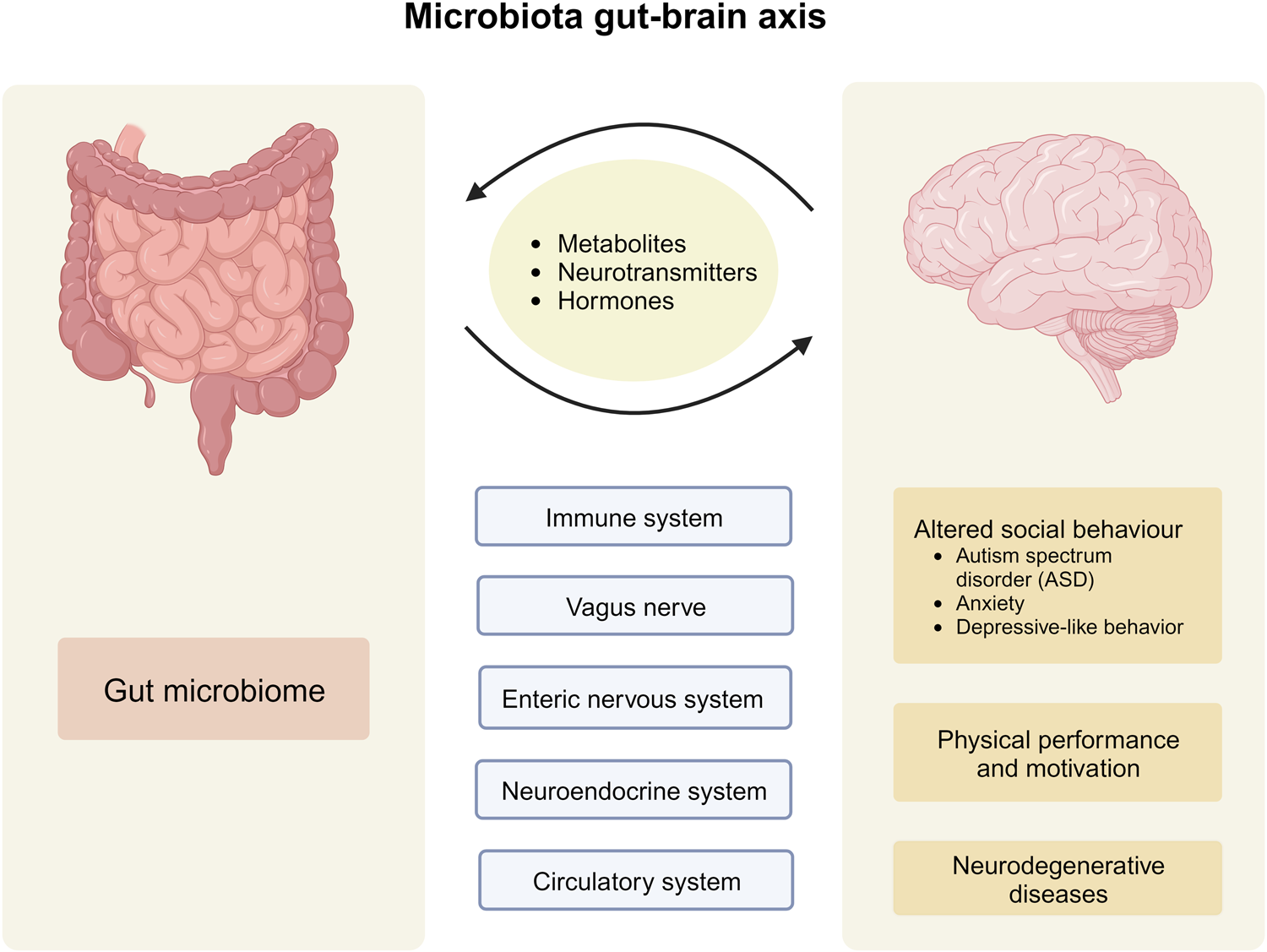
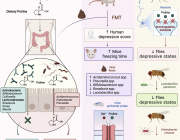
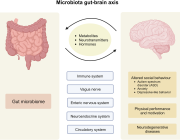
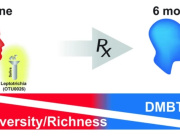
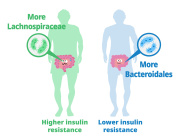
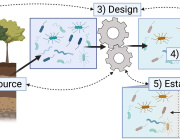
Impressive previous studies by Dr. Aleksandar Kostic along with his team have shown that industrial lifestyles are correlated with both a lower diversity in the gut microbiome and increased incidence of chronic diseases, such as obesity and autoimmune diseases. Examining our ancestral gut microbiome may provide insights into aspects of human–microbiome symbioses that have become altered in the present-day industrialized world. Reconstruction of metagenome-assembled genomes (MAGs) is an emerging approach to recover high-quality genomes and previously undescribed species-level genome bins (SGBs) from shotgun metagenomics data. Sequencing reads are de novo assembled into contiguous sequences (contigs), and contigs are binned to form draft genomes.
To summarize, palaeofaeces with well-preserved DNA are abundant sources of microbial genomes, including previously undescribed microbial species, that may elucidate the evolutionary histories of human microbiomes. Similar future studies tapping into the richness of palaeofaeces will not only expand our knowledge of the human microbiome, but may also lead to the development of approaches to restore present-day gut microbiomes to their ancestral state.
Full article: https://www.nature.com/articles/s41586-021-03532-0#Abs1
This topic will be discussed during the World Congress on Targeting Microbiota 2021 which will be held on October 20-22, 2021.
Targeting Microbiota 2021 Congress
October 20-22, 2021
www.microbiota-site.com